Identifying Indicator Species in Ecological Habitats using Deep Optimal Feature Learning
PLOS ONE,
Yiting Tsai, Susan A. Baldwin, Bhushan Gopaluni
[PDF] [Code]
Abstract
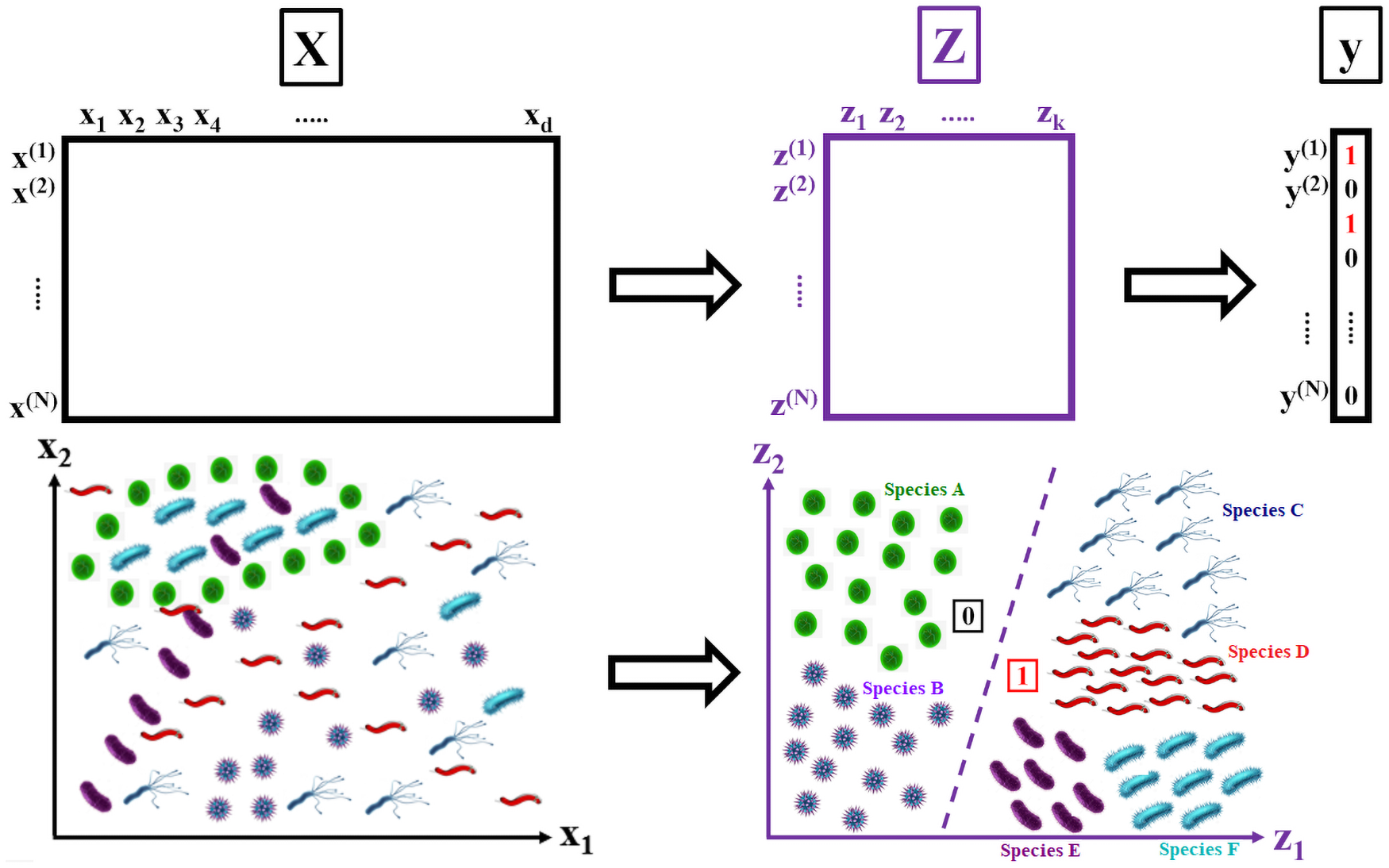
Much of the current research on supervised modelling is focused on maximizing outcome prediction accuracy. However, in engineering disciplines, an arguably more important goal is that of feature extraction, the identification of relevant features associated with the various outcomes. For instance, in microbial communities, the identification of keystone species can often lead to improved prediction of future behavioral shifts. This paper proposes a novel feature extractor based on Deep Learning, which is largely agnostic to underlying assumptions regarding the training data. Starting from a collection of microbial species abundance counts, the Deep Learning model first trains itself to classify the selected distinct habitats. It then identifies indicator species associated with the habitats. The results are then compared and contrasted with those obtained by traditional statistical techniques. The indicator species are similar when compared at top taxonomic levels such as Domain and Phylum, despite visible differences in lower levels such as Class and Order. More importantly, when our estimated indicators are used to predict final habitat labels using simpler models (such as Support Vector Machines and traditional Artificial Neural Networks), the prediction accuracy is improved. Overall, this study serves as a preliminary step that bridges modern, black-box Machine Learning models with traditional, domain expertise-rich techniques.
Read or Download: PDF